Genetic diversity of introduced Chinook salmon in Chile
January 10, 2019Correa C and Moran P. 2017. Polyphyletic ancestry of expanding Patagonian Chinook salmon populations. Nature Scientific Reports 7: 14338.
In a nutshell
- Although multiple independent introduction events of Chinook salmon have occurred in many parts of the world, it is unknown which events have led to self-sustaining populations
- Researchers used genetic markers to identify which of the potential multiple stocks of Chinook salmon introduced to the Patagonia region of Chile became established, and examined the role of genetic diversity in the early stages of biological invasions
- Results suggest that Chilean Chinook salmon are derived from almost 150 North American donor stocks, but a distinct separation of ancestral lineages between northern and southern Patagonian populations was also detected
- Although invasive species are often assumed to exhibit less genetic variation due to genetic bottlenecks and founder effects, genetic diversity in Patagonian populations was remarkably high and resembled that of Chinook in the native range despite the multiple sources of introduction
- Multiple sources of introduced Chinook prevented conclusive determination of the relationship between ancestral diversity and genetic diversity among populations in Chile, supporting the view that invasion success or failure is dependent on complex interactions among numerous genetic, environmental, and anthropogenic factors
Though native to the northern Pacific Ocean, Chinook salmon have been introduced to many other regions of the world over the past century as part of efforts to establish viable commercial and recreational fisheries. The establishment of non-native species in new areas often has negative environmental impacts, and Chinook are no exception – where introduced, Chinook salmon are often considered to be harmful biological invaders because they compete with native species. At the same time, however, introductions of Chinook salmon populations into such regions as the North American Great Lakes, New Zealand, and Chile have also provided scientists with unique opportunities to explore issues of contemporary evolution of a non-native salmonid over large scales and under natural conditions, with the goal of understanding colonization dynamics that also apply throughout their native range.
The Patagonia region of southern Chile is a prominent example of the successful establishment of Chinook populations, but why Chinook thrive in southern Chilean waters and not in other locations where they have been introduced is unknown. Genetic diversity among the introduced populations is one potentially important factor that may have a significant influence on establishment success (or failure) that has received comparatively little attention. On the one hand, genetic diversity may be considerably lower in invasive populations than in populations in the native range because of the relatively small number of individuals initially introduced from a single stock, thereby resulting in a genetic bottleneck; moreover, smaller populations are also more susceptible to genetic drift and founder effects. On the other hand, invasive populations can also be composed of a complex mix (ie “admixture”) of individuals taken from multiple source stocks, which should – at least in theory – result in greater genetic diversity in the relocated populations.
To explore these alternatives, Correa and Moran collected tissue samples from Chinook populations in four discrete basins along approximately 900 km of the coastline of Chilean Patagonia and used modern genetic forensics to identify the ancestral sources of the salmon in this region. In addition, the authors also examined the relationship between the degree of genetic diversity and successful establishment of Chinook salmon populations. Analysis of 13 microsatellite loci indicated that Chinook in southern Patagonia could be traced back almost exclusively to Chinook salmon stocks native to the Lower Columbia River that were introduced into Patagonia in the late 1970s and early 1980s in order to establish hatcheries in which fish are grown until the smolt stage and then released into the ocean to feed. In contrast, Chinook salmon populations in the northern part of the study area, which were primarily introduced via net-pen aquaculture operations in the 1990s, were found to derive from a wide range of sources, including Willamette River, northern coastal Oregon, middle Columbia River, and Puget Sound stocks. The analysis also revealed the presence of genetic contributions from unknown introductions, an indication that not all historical introduction events were documented.
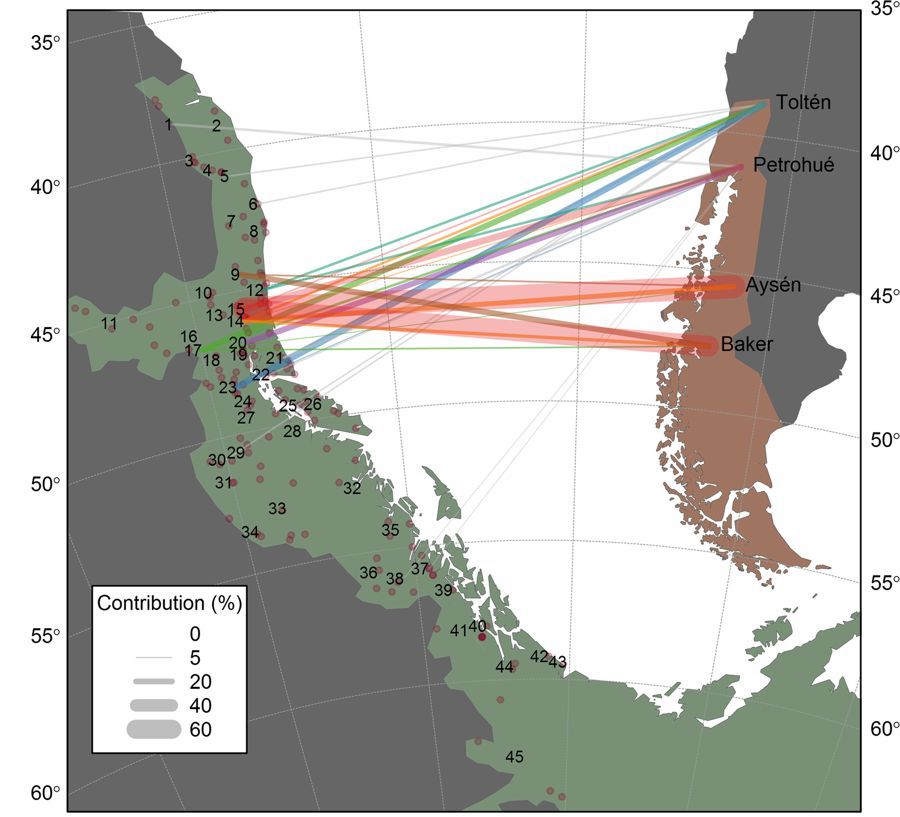
Analysis of the genetic diversity among these Chinook salmon populations revealed two unexpected and surprising results. First, overall diversity among Chilean Chinook salmon was not only higher than expected, but was nearly identical to that of native populations in the northern Pacific, suggesting that no genetic bottlenecks had occurred and undoubtedly reflecting multiple introduction events. Second, Chinook salmon populations in southern Patagonia, which are of highly mixed origin, exhibited no greater degree of allelic heterozygosity than did Chinook salmon populations in northern Patagonia, which derived primarily from a single native stock. In light of these results, the authors concluded that lineage admixture of these populations reduced the effectiveness and accuracy of assignment tests provided by traditional mixture models, and that future models should be viewed with some caution due to this potential limitation.
Correa and Moran’s analysis demonstrates that simple predictions about introductions and genetic diversity may not always be warranted given the complex interactions that can arise from admixture lineages. The success of Chinook salmon in Patagonia, despite various genetic backgrounds, points to the need for further research to uncover the specific aspects of genetic architecture that influences the ability of introduced Chinook salmon populations to adapt and flourish in new areas.
Science Spotlight by Ken Ferguson
